Nucleocytoplasmic Translocations

Nucleocytoplasmic protein translocations in Dictyostelium discoideum. Calmodulin binding proteins are shown in green text. Peptide domains and CaM that are involved in specific events are shown in blue.
Dictyostelium is a eukaryotic microbe in which the nucleoli exist as 2-3 patches at the periphery of the nuclei rather than as a single central nucleolus. Our lab was the first to identify and characterize the nuclear/nucleolar protein. Nucleomorphin (NumA1), which localizes to primarily to nucleoli, was so named because it regulates the number of nuclei in Dictyostelium cells. The gene encoding NumA1 was isolated using the Calmodulin-Binding Overlay Technique (CaMBOT) and the protein was subsequently validated as a calcium-dependent calmodulin (CaM) binding protein (CaMBP). A CaM-Binding Domain or CaMBD (171EDVSRFIKGKLLQKQQKIYKDLERF195) containing both calcium-dependent-binding motifs and an IQ-like motif for calcium-independent binding has been experimentally validated. A nuclear/nucleolar localization sequence was also revealed (48KKSYQDPEIIAHSRPRK64) that includes both a putative bipartite and classical NLS. Four functional Nucleolar Localization Sequences (NoLS) rich in basic residues (e.g., RPRK) were identified in NumA1 and shown to translocated FITC to nucleoli. NumA1 is regulated by the stalk cell inducing morphogen DIF-1.
NuMA1, a member of the nucleoplasmin family, contains a glu/asp inverted repeat or DEED sequence which when deleted results in multinuclearity. Database searches indicated that NumA1 is a member of the BRCT-domain family of cell cycle proteins and continued research would strengthen this cell cycle link.
NumA1 Binds to CBP4a and Puromycin Sensitive Aminopeptidase
Using a Yeast 2 Hybrid Screen, we showed that NumA1 binds to the calcium-binding protein Cbp4a in the nucleolus in a calcium-dependent manner. This interaction requires residues found in the DEED sequence. In contrast puromycin-sensitive aminopeptidase (PsaA) also bound to NumA1 but this association was limited to the nucleoplasm.
Puromycin Sensitive Aminopeptidase Binds to Cdk5
PsaA and Cdk5 (cyclin-dependent kinase 5) coimmunoprecipitate in CaM-pulldowns. The three proteins were also shown to reside in isolated nuclei showing for the first time that CaM is nuclear in Dictyostelium. The CaMBDs of PsaA and Cdk5 were identified and deletion of the CaMBD of Cdk5 led to increased nuclear retention indicating that CaM mediates the nucleocytoplasmic transport of this protein.
In addition to NumA1 and CBP4a which localize throughout nucleoli, FhkA (a Rad53 homolog) was found to be a nucleolar protein that localizes to the nucleolar periphery. Cbp4a, PsaA, Cdk5 and FhkA show distinct nucleoplasmic translocations during mitosis as shown in the above figure. Here we show the immunolocalizations for Cbp4a and FhkA.
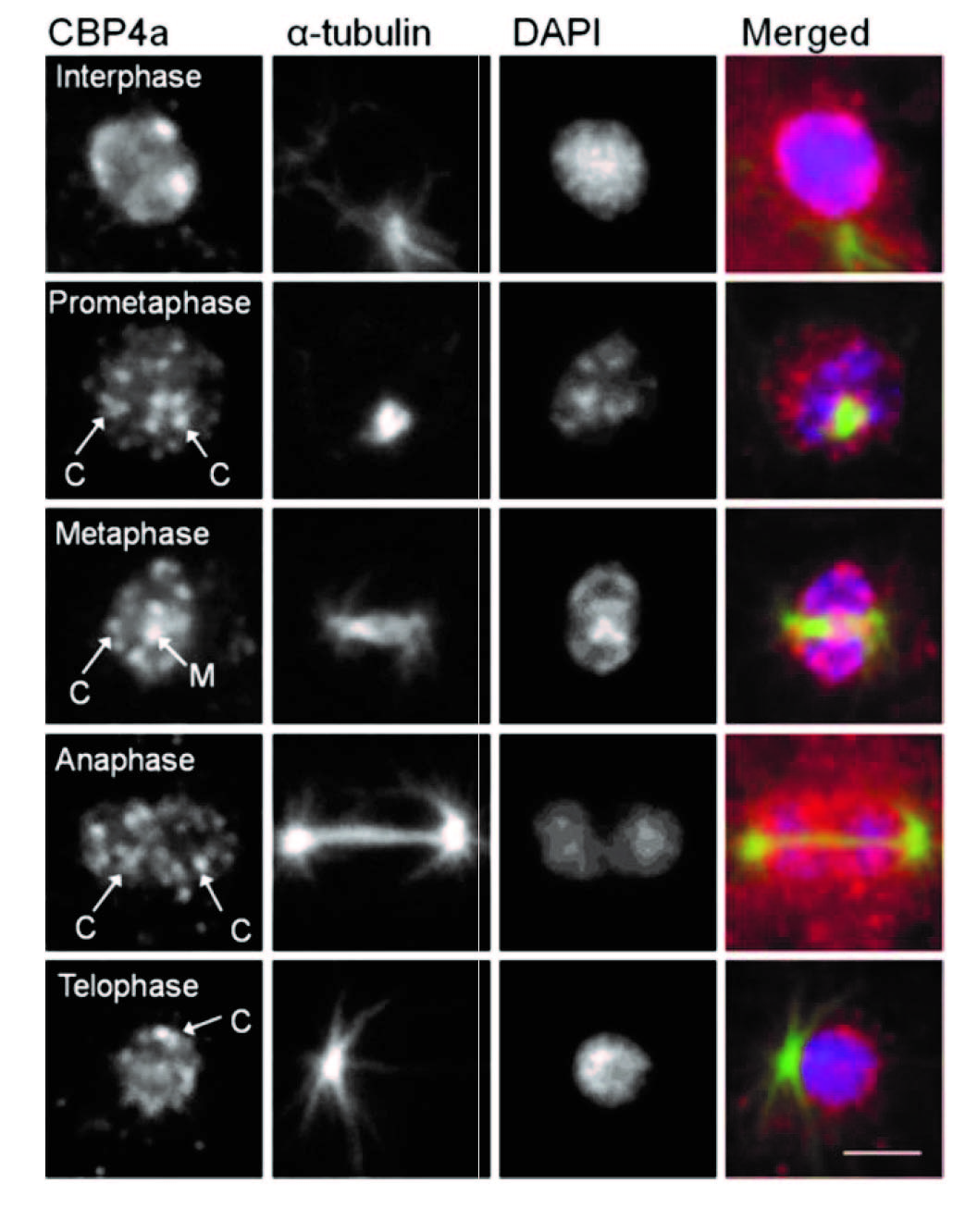
Figure. Immunolocalization of Cbp4a during mitosis.
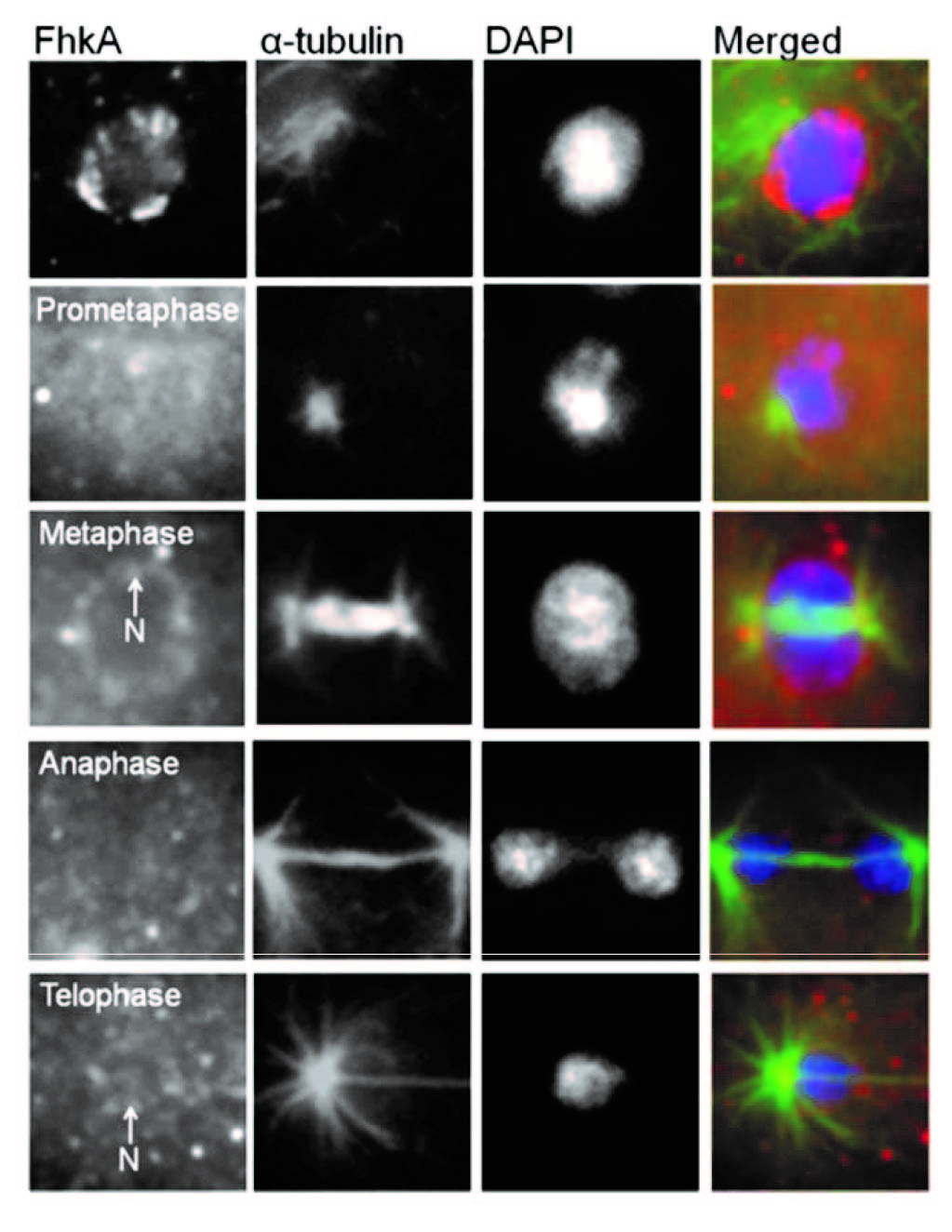
Figure. Immunolocalization of FhkA during mitosis.
The study of NumA1 and its binding partners and their associates is the most detailed for a nucleolar protein and for any nuclear protein. Considering the relationship to the cell cycle, nucleolar/nucleolar targeting and calmodulin binding, this protein presents an interesting subject for continued analyses.
More about Andrew Catalano, PhD, Smart-Biology, Biology Animations:
https://www.smart-biology.com/
More about Robert Huber, Asst. Professor, Biology, Trent University:
http://huberlab.weebly.com/
More about Michael Myre, Asst. Professor, Biology, University of Massachusetts, Lowell:
https://www.uml.edu/Sciences/biology/faculty/Myre-Michael.aspx
References
Myre, Michael A. and Danton H. O'Day, 2002. Nucleomorphin: A novel, acidic, nuclear calmodulin-binding protein from Dictyostelium that regulates nuclear number. J. Biol. Chem. 277: 19735-19744.
O’Day, Danton H., 2003. CaMBOT: Profiling and Characterizing Calmodulin Binding Proteins. Cellular Signalling 15:347-355.
Myre, Michael A. and Danton H. O’Day, 2004. Dictyostelium calcium-binding protein 4a interacts with Nucleomorphin, a BRCT-domain protein that regulates nuclear number. Biochemical Biophysical Research Communications 322: 665-671.
Myre, M.A. and Danton H. O’Day, 2004. Dictyostelium nucleomorphin is a member of the BRCT-domain family of cell cycle checkpoint proteins. Biochimica Biophysica Acta, General Subjects.1675: 192-197.
Myre, M.A. and Danton H. O’Day, 2005. An N-terminal nuclear localization sequence but not the calmodulin-binding domain mediates nuclear localization of nucleomorphin, a protein that regulates nuclear number in Dictyostelium.Biochemical Biophysical Research Communications 332:157-166.
Catalano, A. and D.H. O’Day, 2008. Calmodulin-binding proteins in the model organism Dictyostelium: A complete and critical review. Cellular Signalling 20: 277-291.
O’Day, Danton H., Yekaterina Poloz and Michael A. Myre, 2009. Differentiation inducing factor-1 induces gene and protein expression of the Dictyostelium nuclear calmodulin-binding protein nucleomorphin. Cellular Signalling 21: 317-323.
Catalano, Andrew and Danton H. O’Day, 2011. Nucleolar localization during mitosis and identification of nuclear/nucleolar localization signals of the calmodulin-binding protein nucleomorphin during growth and mitosis in Dictyostelium. Histochem. Cell Biol. 135: 239-249.
Robert J. Huber and Danton H. O’Day, 2011. Nucleocytoplasmic transfer of cyclin dependent kinase 5 and its binding to puromycin-sensitive aminopeptidase in Dictyostelium discoideum. Histochem Cell Biology 136: 177-189.
Andrew Catalano, Yekaterina Poloz and Danton H. O’Day, 2011. Dictyostelium puromycin-sensitive aminopeptidase A is a nucleoplasmic nucleomorphin-binding protein that relocates to the cytoplasm during mitosis. Histochem Cell Biol 136: 677–688
Yekaterina Poloz, Andrew Catalano and Danton H. O’Day, 2012. Bestatin inhibits cell growth, division and differentiation in Dictyostelium through inhibition of puromycin sensitive aminopeptidase. Eukaryotic Cell 11: 545-557
Danton H. O’Day and Aldona Budniak, 2015. Nucleocytoplasmic Protein Translocation during Mitosis in the Social Amoebozoan Dictyostelium discoideum. Biological Reviews 90: 126-141
All information is the property of Danton H. O'Day, 2017© and may be used by individuals without prior permission if proper credit is given. If you would like to distribute or communicate this information to others in any form other than citing this website, you need to obtain permission.
Last update: July 2017.